from oauthlib.oauth2 import BackendApplicationClient
from requests_oauthlib import OAuth2Session
client_id = "your-client-id"
client_secret = "your-client-secret"
client = BackendApplicationClient(client_id=client_id)
oauth = OAuth2Session(client=client)
token = oauth.fetch_token(
token_url="https://identity.dataspace.copernicus.eu/auth/realms/CDSE/protocol/openid-connect/token",
client_secret=client_secret,
include_client_id=True,
)Data acquisition
1 Sentinel-2 Images
This section describes how Sentinel-2 images are retrieved, visualised, and organised for model training.
1.1 What is Sentinel-2?
Sentinel-2 is an Earth observation mission from the European Space Agency (ESA), part of the Copernicus programme. It consists of two twin satellites — Sentinel-2A and Sentinel-2B — placed in the same sun-synchronous orbit but phased 180 degrees apart, giving a 5-day revisit time at the equator.
Key characteristics:
- 13 spectral bands spanning visible, near-infrared (NIR) and short-wave infrared (SWIR)
- Spatial resolution of 10 m, 20 m or 60 m depending on the band
- 290 km swath width
- Free and open data policy
These properties make Sentinel-2 a reference dataset for land-cover mapping, agricultural monitoring, forestry, urban change detection and many other applications.
1.1.1 Spectral bands
| Band | Central wavelength (nm) | Resolution (m) | Description |
|---|---|---|---|
| B1 | 443 | 60 | Coastal aerosol |
| B2 | 490 | 10 | Blue |
| B3 | 560 | 10 | Green |
| B4 | 665 | 10 | Red |
| B5 | 705 | 20 | Vegetation red edge |
| B6 | 740 | 20 | Vegetation red edge |
| B7 | 783 | 20 | Vegetation red edge |
| B8 | 842 | 10 | NIR |
| B8A | 865 | 20 | Narrow NIR |
| B9 | 945 | 60 | Water vapour |
| B10 | 1375 | 60 | SWIR – Cirrus |
| B11 | 1610 | 20 | SWIR |
| B12 | 2190 | 20 | SWIR |
1.2 Copernicus Data Space Ecosystem (CDSE) — Sentinel Hub API
The official programmatic access point for Sentinel-2 data is the Sentinel Hub API, part of the Copernicus Data Space Ecosystem (CDSE). It provides two main endpoints:
- Catalog API — search for available Sentinel-2 products by area, date, and cloud cover
- Processing API — download imagery on-the-fly with custom band combinations and formats
To use the Sentinel Hub API you need a free CDSE account — register at https://dataspace.copernicus.eu/. Then, register an OAuth client in the CDSE dashboard to obtain a Client ID and Client Secret.
1.2.1 Authentication
Sentinel Hub uses OAuth2 client credentials. Authenticate with oauthlib and requests_oauthlib:
1.2.2 Searching the catalogue
Use the Catalog API to find Sentinel-2 L2A products for a given area and date range:
catalog_url = "https://sh.dataspace.copernicus.eu/api/v1/catalog/1.0.0/search"
search_body = {
"collections": ["sentinel-2-l2a"],
"datetime": "2021-06-01T00:00:00Z/2021-06-30T23:59:59Z",
"bbox": [23.0, 42.0, 24.0, 43.0], # [west, south, east, north] in WGS84
"limit": 10,
"filter": "eo:cloud_cover < 20",
"filter-lang": "cql2-text",
}
response = oauth.post(catalog_url, json=search_body)
results = response.json()
print(f"Found {len(results['features'])} products")1.2.3 Downloading imagery
Use the Processing API to download a true-colour Sentinel-2 image as a GeoTIFF:
process_url = "https://sh.dataspace.copernicus.eu/api/v1/process"
evalscript = """
//VERSION=3
function setup() {
return { input: ["B04", "B03", "B02"], output: { bands: 3, sampleType: "UINT16" } };
}
function evaluatePixel(sample) {
return [sample.B04, sample.B03, sample.B02];
}
"""
request_body = {
"input": {
"bounds": {
"bbox": [23.0, 42.0, 24.0, 43.0],
"properties": {"crs": "http://www.opengis.net/def/crs/EPSG/0/4326"},
},
"data": [
{
"type": "sentinel-2-l2a",
"dataFilter": {
"timeRange": {
"from": "2021-06-01T00:00:00Z",
"to": "2021-06-30T23:59:59Z",
},
"maxCloudCoverage": 20,
},
}
],
},
"output": {
"width": 512,
"height": 512,
"responses": [{"identifier": "default", "format": {"type": "image/tiff"}}],
},
"evalscript": evalscript,
}
response = oauth.post(process_url, json=request_body)
with open("output.tif", "wb") as f:
f.write(response.content)collections: use"sentinel-2-l2a"for Level-2A products (surface reflectance, atmospherically corrected).bbox: bounding box as[west, south, east, north]in WGS84 (EPSG:4326).filter: CQL2 expression for metadata filtering, e.g."eo:cloud_cover < 20"keeps only scenes with under 20 % cloud cover.
Goal: Use the Sentinel Hub Catalog API to search for Sentinel-2 L2A products over Bulgaria during June 2021 with less than 10 % cloud cover.
Steps:
- Authenticate with OAuth2 using
OAuth2Session+BackendApplicationClientagainst the CDSE token endpoint - Build a Catalog API request body with
collections: ["sentinel-2-l2a"],datetime,bboxcovering Bulgaria, and a cloud-cover filter - POST to the Catalog API endpoint and parse the JSON response
- Print the number of features found
from oauthlib.oauth2 import BackendApplicationClient
from requests_oauthlib import OAuth2Session
# Step 1: Authenticate with OAuth2
client_id = ___ # TODO: your CDSE OAuth client ID
client_secret = ___ # TODO: your CDSE OAuth client secret
client = BackendApplicationClient(client_id=client_id)
oauth = OAuth2Session(client=client)
token = oauth.fetch_token(
token_url=___, # TODO: CDSE token URL
client_secret=client_secret,
include_client_id=True,
)
# Step 2: Build the Catalog API request body
catalog_url = "https://sh.dataspace.copernicus.eu/api/v1/catalog/1.0.0/search"
search_body = {
"collections": ___, # TODO: ["sentinel-2-l2a"]
"datetime": ___, # TODO: "2021-06-01T00:00:00Z/2021-06-30T23:59:59Z"
"bbox": ___, # TODO: Bulgaria bounding box [west, south, east, north]
"limit": 50,
"filter": ___, # TODO: CQL2 cloud cover filter
"filter-lang": "cql2-text",
}
# Step 3: POST to the Catalog API
response = ___ # TODO: oauth.post(catalog_url, json=search_body)
results = response.json()
# Step 4: Print the number of features
print(f"Found {len(results['features'])} products")- The CDSE token URL is
"https://identity.dataspace.copernicus.eu/auth/realms/CDSE/protocol/openid-connect/token". - Bulgaria’s approximate bounding box in WGS84 is
[22.0, 41.0, 29.0, 44.5]. - Use
"eo:cloud_cover < 10"as the CQL2 filter for cloud cover under 10 %. - The
datetimerange format is"start/end"in ISO 8601, e.g."2021-06-01T00:00:00Z/2021-06-30T23:59:59Z".
from oauthlib.oauth2 import BackendApplicationClient
from requests_oauthlib import OAuth2Session
client_id = "your-client-id"
client_secret = "your-client-secret"
client = BackendApplicationClient(client_id=client_id)
oauth = OAuth2Session(client=client)
token = oauth.fetch_token(
token_url="https://identity.dataspace.copernicus.eu/auth/realms/CDSE/protocol/openid-connect/token",
client_secret=client_secret,
include_client_id=True,
)
catalog_url = "https://sh.dataspace.copernicus.eu/api/v1/catalog/1.0.0/search"
search_body = {
"collections": ["sentinel-2-l2a"],
"datetime": "2021-06-01T00:00:00Z/2021-06-30T23:59:59Z",
"bbox": [22.0, 41.0, 29.0, 44.5],
"limit": 50,
"filter": "eo:cloud_cover < 10",
"filter-lang": "cql2-text",
}
response = oauth.post(catalog_url, json=search_body)
results = response.json()
print(f"Found {len(results['features'])} products")Once you have searched the catalogue, you can use the Processing API to download imagery directly, then upload it to S3 so it can be shared with your team. The SSP Cloud provides an S3-compatible object store accessible via s3fs. Here is how to initialise an S3FileSystem and upload a file:
from utils import get_file_system
fs = get_file_system()
# Upload a local file to S3
fs.put("./output.tif", "s3://my-bucket/sentinel2/output.tif")Goal: Use the Sentinel Hub Processing API to download a true-colour Sentinel-2 image for Bulgaria (June 2021) and upload it to your personal S3 bucket.
Steps:
- Authenticate with OAuth2 (reuse the session from Exercise 1)
- Build a Processing API request with bbox, time range, evalscript for true-colour RGB, and output as
image/tiff - POST to the Processing API endpoint and save the response content locally
- Initialise an
S3FileSystemwithget_file_system()and upload the file withfs.put()
from utils import get_file_system
# Step 1: Reuse the authenticated OAuth2 session from Exercise 1
# Step 2: Build the Processing API request
process_url = "https://sh.dataspace.copernicus.eu/api/v1/process"
evalscript = """
//VERSION=3
function setup() {
return { input: ["B04", "B03", "B02"], output: { bands: 3, sampleType: "UINT16" } };
}
function evaluatePixel(sample) {
return [sample.B04, sample.B03, sample.B02];
}
"""
request_body = {
"input": {
"bounds": {
"bbox": ___, # TODO: Bulgaria bounding box [west, south, east, north]
"properties": {"crs": "http://www.opengis.net/def/crs/EPSG/0/4326"},
},
"data": [
{
"type": ___, # TODO: "sentinel-2-l2a"
"dataFilter": {
"timeRange": {
"from": ___, # TODO: start date ISO 8601
"to": ___, # TODO: end date ISO 8601
},
"maxCloudCoverage": ___, # TODO: max cloud cover percentage
},
}
],
},
"output": {
"width": 512,
"height": 512,
"responses": [{"identifier": "default", "format": {"type": ___}}], # TODO: "image/tiff"
},
"evalscript": evalscript,
}
# Step 3: POST and save locally
response = ___ # TODO: oauth.post(process_url, json=request_body)
local_path = "bulgaria_rgb.tif"
with open(local_path, "wb") as f:
f.write(___) # TODO: response.content
# Step 4: Upload to S3
fs = ___ # TODO: get_file_system()
fs.put(___, ___) # TODO: local_path, S3 destination path- The Processing API URL is
"https://sh.dataspace.copernicus.eu/api/v1/process". - Use
[22.0, 41.0, 29.0, 44.5]for the Bulgaria bounding box. - Time range uses ISO 8601:
"2021-06-01T00:00:00Z"to"2021-06-30T23:59:59Z". - Set
maxCloudCoverageto10(integer, not a string range). - The response content (
response.content) is the raw TIFF bytes — write them to a file in binary mode. get_file_system()reads SSP Cloud credentials from environment variables.
from utils import get_file_system
process_url = "https://sh.dataspace.copernicus.eu/api/v1/process"
evalscript = """
//VERSION=3
function setup() {
return { input: ["B04", "B03", "B02"], output: { bands: 3, sampleType: "UINT16" } };
}
function evaluatePixel(sample) {
return [sample.B04, sample.B03, sample.B02];
}
"""
request_body = {
"input": {
"bounds": {
"bbox": [22.0, 41.0, 29.0, 44.5],
"properties": {"crs": "http://www.opengis.net/def/crs/EPSG/0/4326"},
},
"data": [
{
"type": "sentinel-2-l2a",
"dataFilter": {
"timeRange": {
"from": "2021-06-01T00:00:00Z",
"to": "2021-06-30T23:59:59Z",
},
"maxCloudCoverage": 10,
},
}
],
},
"output": {
"width": 512,
"height": 512,
"responses": [{"identifier": "default", "format": {"type": "image/tiff"}}],
},
"evalscript": evalscript,
}
response = oauth.post(process_url, json=request_body)
local_path = "bulgaria_rgb.tif"
with open(local_path, "wb") as f:
f.write(response.content)
fs = get_file_system()
fs.put(local_path, "s3://my-bucket/sentinel2/bulgaria_rgb.tif")1.3 What is rasterio?
rasterio is a Python library built on top of GDAL for reading and writing geospatial raster data (GeoTIFF, etc.). It is the standard tool for working with satellite imagery in Python.
Key concepts:
- Raster = a grid of pixels organised into bands (layers). Each band represents one channel of data (e.g. a spectral band from a satellite sensor).
rasterio.open(path)returns aDatasetReader— works with local files and HTTPS URLs alike.src.read(bands)returns a NumPy array with shape(bands, H, W). Band indices are 1-based: band 1 is the first band, not band 0.src.crs— the coordinate reference system attached to the raster.src.bounds— the geographic extent as(left, bottom, right, top).src.profile/src.meta— full metadata dictionary (driver, dtype, nodata value, affine transform, etc.).
Why band indexing matters: Sentinel-2 images have 13 spectral bands (see the table above). Reading specific band combinations reveals different features:
src.read([4, 3, 2])→ true-colour RGB (Red, Green, Blue)src.read([8, 4, 3])→ false-colour composite (NIR, Red, Green) — vegetation appears bright redsrc.read([8, 4])→ bands needed for NDVI (Normalised Difference Vegetation Index)
You will practise these combinations in the exercises below.
2 Accessing pre-processed patches
For this project, pre-processed Sentinel-2 patches are publicly available on the SSP Cloud object storage. You can access them directly over HTTPS — no authentication or special tools required.
The URL follows this pattern:
https://minio.lab.sspcloud.fr/projet-hackathon-ntts-2025/data-preprocessed/patchs/CLCplus-Backbone/SENTINEL2/{NUTS}/{year}/{size}/{patch_id}.tifEach pre-processed patch is a GeoTIFF covering a 250 x 250 pixel area (2 500 m at 10 m resolution). The URL path encodes the key metadata:
.../SENTINEL2/{NUTS}/{year}/{size}/{xmin}_{ymin}_{idx}_{id}.tif| Field | Example value | Meaning |
|---|---|---|
| NUTS | FRJ27 |
NUTS-3 region (Haute-Garonne, near Toulouse) |
| year | 2018 |
Acquisition year |
| size | 250 |
Tile size in pixels |
| xmin, ymin | 3649890, 2331750 |
Lower-left corner in EPSG:3035 (metres) |
rasterio can open these URLs directly, just like local files:
import rasterio
import numpy as np
tile_url = (
"https://minio.lab.sspcloud.fr/projet-hackathon-ntts-2025/"
"data-preprocessed/patchs/CLCplus-Backbone/SENTINEL2/"
"FRJ27/2018/250/3649890_2331750_0_937.tif"
)
with rasterio.open(tile_url) as src:
tile_crs = src.crs
tile_bounds = src.bounds
tile_count = src.count
tile_height = src.height
tile_width = src.width
# Read RGB bands: B4 (Red), B3 (Green), B2 (Blue)
rgb_data = src.read([4, 3, 2])
print(f"CRS: {tile_crs}")
print(f"Bounds: {tile_bounds}")
print(f"Shape: {tile_count} bands x {tile_height} x {tile_width} px")CRS: EPSG:3035
Bounds: BoundingBox(left=3649890.0, bottom=2331750.0, right=3652390.0, top=2334250.0)
Shape: 14 bands x 250 x 250 pxNow let’s display the tile as an RGB composite (true-colour: Red, Green, Blue):
import matplotlib.pyplot as plt
# Transpose to (H, W, 3) and normalize for display
rgb = np.transpose(rgb_data, (1, 2, 0)).astype(np.float32)
p98 = np.percentile(rgb, 98)
rgb = np.clip(rgb / p98, 0, 1)
fig, ax = plt.subplots(figsize=(5, 5))
ax.imshow(rgb)
ax.set_title("Sentinel-2 RGB composite (B4, B3, B2) — FRJ27")
ax.axis("off")
plt.tight_layout()
plt.show()
Goal: Open a different pre-processed Sentinel-2 patch via its HTTPS URL, extract the RGB bands, normalize the image, and display it.
Steps:
- Open the tile with
rasterio.open(tile_url)and read bands 4, 3, 2 (Red, Green, Blue) - Store the CRS and bounds from the rasterio source
- Transpose the array to
(H, W, 3)and normalize for display (divide by the 98th percentile, clip to[0, 1]) - Display the RGB composite with
matplotlib
import rasterio
import numpy as np
import matplotlib.pyplot as plt
tile_url = (
"https://minio.lab.sspcloud.fr/projet-hackathon-ntts-2025/"
"data-preprocessed/patchs/CLCplus-Backbone/SENTINEL2/"
"FRJ27/2018/250/3649890_2331750_0_937.tif"
)
# Step 1: Open the tile and read RGB bands (4, 3, 2)
with rasterio.open(tile_url) as src:
rgb_data = ___ # TODO: read bands 4, 3, 2
tile_crs = ___ # TODO
tile_bounds = ___ # TODO
# Step 2: Transpose to (H, W, 3) and normalize
rgb = ___ # TODO: np.transpose, then normalize with 98th percentile and clip
# Step 3: Display
fig, ax = plt.subplots(figsize=(5, 5))
___ # TODO: ax.imshow(...)
ax.set_title("Sentinel-2 RGB composite")
ax.axis("off")
plt.tight_layout()
plt.show()- Inside the
with rasterio.open(tile_url) as src:block, usesrc.read([4, 3, 2])to get a(3, H, W)array. src.crsandsrc.boundsgive you the CRS and bounding box.- Transpose with
np.transpose(rgb_data, (1, 2, 0)), cast tofloat32, then normalize: divide bynp.percentile(rgb, 98)and clip to[0, 1]withnp.clip.
import rasterio
import numpy as np
import matplotlib.pyplot as plt
tile_url = (
"https://minio.lab.sspcloud.fr/projet-hackathon-ntts-2025/"
"data-preprocessed/patchs/CLCplus-Backbone/SENTINEL2/"
"FRJ27/2018/250/3649890_2331750_0_937.tif"
)
with rasterio.open(tile_url) as src:
rgb_data = src.read([4, 3, 2])
tile_crs = src.crs
tile_bounds = src.bounds
rgb = np.transpose(rgb_data, (1, 2, 0)).astype(np.float32)
p98 = np.percentile(rgb, 98)
rgb = np.clip(rgb / p98, 0, 1)
fig, ax = plt.subplots(figsize=(5, 5))
ax.imshow(rgb)
ax.set_title("Sentinel-2 RGB composite")
ax.axis("off")
plt.tight_layout()
plt.show()Goal: Inspect the raster metadata of a Sentinel-2 tile using src.profile, then create a false-colour composite using bands 8 (NIR), 4 (Red), 3 (Green). In this composite, vegetation appears bright red because it reflects strongly in the near-infrared.
Steps:
- Open the tile with
rasterio.open(tile_url)and printsrc.profile - Read bands
[8, 4, 3](NIR, Red, Green) - Normalize the false-colour image (same method as Exercise 3: 98th percentile + clip)
- Display the true-colour RGB (from Exercise 3) and false-colour composite side by side
import rasterio
import numpy as np
import matplotlib.pyplot as plt
tile_url = (
"https://minio.lab.sspcloud.fr/projet-hackathon-ntts-2025/"
"data-preprocessed/patchs/CLCplus-Backbone/SENTINEL2/"
"FRJ27/2018/250/3649890_2331750_0_937.tif"
)
# Step 4a: Open the tile and print the raster profile
with rasterio.open(tile_url) as src:
print(___) # TODO: print src.profile
# Step 4b: Read false-colour bands (NIR=8, Red=4, Green=3)
fc_data = ___ # TODO: src.read([8, 4, 3])
# Step 4c: Normalize for display
fc = np.transpose(fc_data, (1, 2, 0)).astype(np.float32)
p98 = ___ # TODO: np.percentile(fc, 98)
fc = ___ # TODO: np.clip(fc / p98, 0, 1)
# Step 4d: Display side by side with the true-colour RGB
fig, axes = plt.subplots(1, 2, figsize=(10, 5))
axes[0].imshow(rgb)
axes[0].set_title("True colour (B4, B3, B2)")
axes[0].axis("off")
___ # TODO: axes[1].imshow(fc)
axes[1].set_title("False colour (B8, B4, B3)")
axes[1].axis("off")
plt.tight_layout()
plt.show()src.profileis a dictionary — justprint(src.profile)to see driver, dtype, crs, transform, etc.src.read([8, 4, 3])reads NIR, Red, Green in that order → shape(3, H, W).- Normalize exactly as in Exercise 3: transpose to
(H, W, 3), divide bynp.percentile(fc, 98), clip to[0, 1].
import rasterio
import numpy as np
import matplotlib.pyplot as plt
tile_url = (
"https://minio.lab.sspcloud.fr/projet-hackathon-ntts-2025/"
"data-preprocessed/patchs/CLCplus-Backbone/SENTINEL2/"
"FRJ27/2018/250/3649890_2331750_0_937.tif"
)
with rasterio.open(tile_url) as src:
print(src.profile)
fc_data = src.read([8, 4, 3])
fc = np.transpose(fc_data, (1, 2, 0)).astype(np.float32)
p98 = np.percentile(fc, 98)
fc = np.clip(fc / p98, 0, 1)
fig, axes = plt.subplots(1, 2, figsize=(10, 5))
axes[0].imshow(rgb)
axes[0].set_title("True colour (B4, B3, B2)")
axes[0].axis("off")
axes[1].imshow(fc)
axes[1].set_title("False colour (B8, B4, B3)")
axes[1].axis("off")
plt.tight_layout()
plt.show()Goal: Compute the Normalised Difference Vegetation Index (NDVI) from a Sentinel-2 tile and display it as a colour-coded map.
NDVI is defined as:
\[ \text{NDVI} = \frac{\text{NIR} - \text{Red}}{\text{NIR} + \text{Red}} \]
NDVI values range from −1 to +1:
- Near +1 → dense, healthy vegetation
- Near 0 → bare soil, urban areas
- Negative → water bodies
Steps:
- Read bands 8 (NIR) and 4 (Red) as
float32 - Compute NDVI pixel-wise, handling division by zero
- Display with
imshow(cmap="RdYlGn", vmin=-1, vmax=1)and a colorbar
import rasterio
import numpy as np
import matplotlib.pyplot as plt
tile_url = (
"https://minio.lab.sspcloud.fr/projet-hackathon-ntts-2025/"
"data-preprocessed/patchs/CLCplus-Backbone/SENTINEL2/"
"FRJ27/2018/250/3649890_2331750_0_937.tif"
)
# Step 5a: Read NIR (band 8) and Red (band 4) as float32
with rasterio.open(tile_url) as src:
nir = ___ # TODO: src.read(8).astype(np.float32)
red = ___ # TODO: src.read(4).astype(np.float32)
# Step 5b: Compute NDVI (handle division by zero)
ndvi = ___ # TODO: np.where(nir + red == 0, 0, (nir - red) / (nir + red))
# Step 5c: Display
fig, ax = plt.subplots(figsize=(6, 5))
im = ___ # TODO: ax.imshow(ndvi, cmap="RdYlGn", vmin=-1, vmax=1)
ax.set_title("NDVI — FRJ27 (2018)")
ax.axis("off")
___ # TODO: fig.colorbar(im, ax=ax, shrink=0.8, label="NDVI")
plt.tight_layout()
plt.show()src.read(8)reads a single band → shape(H, W). Cast tofloat32for division.- Use
np.where(nir + red == 0, 0, (nir - red) / (nir + red))to avoid division by zero. cmap="RdYlGn"gives a red-yellow-green colour scale: red for low NDVI, green for high.fig.colorbar(im, ax=ax, shrink=0.8, label="NDVI")adds a colour legend.
import rasterio
import numpy as np
import matplotlib.pyplot as plt
tile_url = (
"https://minio.lab.sspcloud.fr/projet-hackathon-ntts-2025/"
"data-preprocessed/patchs/CLCplus-Backbone/SENTINEL2/"
"FRJ27/2018/250/3649890_2331750_0_937.tif"
)
with rasterio.open(tile_url) as src:
nir = src.read(8).astype(np.float32)
red = src.read(4).astype(np.float32)
ndvi = np.where(nir + red == 0, 0, (nir - red) / (nir + red))
fig, ax = plt.subplots(figsize=(6, 5))
im = ax.imshow(ndvi, cmap="RdYlGn", vmin=-1, vmax=1)
ax.set_title("NDVI — FRJ27 (2018)")
ax.axis("off")
fig.colorbar(im, ax=ax, shrink=0.8, label="NDVI")
plt.tight_layout()
plt.show()3 Geo-manipulation with GeoPandas
Satellite imagery is inherently geospatial. Before diving into machine-learning pipelines, it is important to know how to manipulate geographic data. GeoPandas extends pandas with geometry support, making CRS transforms, spatial joins, and plotting straightforward.
3.1 Coordinate Reference Systems
Every raster and vector dataset is tied to a coordinate reference system (CRS) that defines how coordinates map to locations on Earth. The two CRS you will encounter most often in this project are:
| CRS | EPSG Code | Units | Use case |
|---|---|---|---|
| WGS 84 | 4326 | Degrees (lat/lon) | GPS, web maps, bounding-box queries |
| ETRS89-LAEA | 3035 | Metres | European statistical grids, CLC+ patches |
Converting between them is a one-liner with GeoPandas:
import geopandas as gpd
from shapely.geometry import box
# Create a GeoDataFrame with the tile extent in EPSG:3035
tile_geom = box(*tile_bounds)
gdf = gpd.GeoDataFrame({"tile": ["FRJ27"]}, geometry=[tile_geom], crs="EPSG:3035")
print("EPSG:3035 bounds:")
print(gdf.total_bounds)
# Convert to WGS84 (latitude/longitude)
gdf_wgs84 = gdf.to_crs("EPSG:4326")
print("\nEPSG:4326 bounds:")
print(gdf_wgs84.total_bounds)EPSG:3035 bounds:
[3649890. 2331750. 3652390. 2334250.]
EPSG:4326 bounds:
[ 1.66386725 43.75893034 1.6979368 43.78382633]3.2 Building geometries from tile metadata
The patch filename encodes the lower-left corner coordinates. Combined with the tile size (2 500 m), you can reconstruct the bounding box and create a shapely geometry:
patch_filename = "3649890_2331750_0_937.tif"
parts = patch_filename.replace(".tif", "").split("_")
xmin, ymin = int(parts[0]), int(parts[1])
tile_size_m = 2500 # 250 px x 10 m/px
xmax = xmin + tile_size_m
ymax = ymin + tile_size_m
tile_box = box(xmin, ymin, xmax, ymax)
print(f"Bounding box (EPSG:3035): [{xmin}, {ymin}, {xmax}, {ymax}]")
print(f"Shapely geometry: {tile_box}")Bounding box (EPSG:3035): [3649890, 2331750, 3652390, 2334250]
Shapely geometry: POLYGON ((3652390 2331750, 3652390 2334250, 3649890 2334250, 3649890 2331750, 3652390 2331750))3.3 Working with GeoDataFrames
In practice, you can build a GeoDataFrame directly from rasterio metadata — no filename parsing needed:
tile_gdf = gpd.GeoDataFrame(
{"tile": ["FRJ27"], "year": [2018]},
geometry=[box(*tile_bounds)],
crs=tile_crs,
)
tile_gdf| tile | year | geometry | |
|---|---|---|---|
| 0 | FRJ27 | 2018 | POLYGON ((3652390 2331750, 3652390 2334250, 36... |
3.4 Overlaying boundaries on images
Combining a raster image with vector boundaries is a common operation. Use imshow with the extent parameter to position the image in coordinate space, then overlay the GeoDataFrame boundary:
fig, ax = plt.subplots(figsize=(6, 6))
extent = [tile_bounds.left, tile_bounds.right, tile_bounds.bottom, tile_bounds.top]
ax.imshow(rgb, extent=extent)
tile_gdf.boundary.plot(ax=ax, color="red", linewidth=2)
ax.set_xlabel("Easting (m)")
ax.set_ylabel("Northing (m)")
ax.set_title("Sentinel-2 tile with boundary overlay (EPSG:3035)")
plt.tight_layout()
plt.show()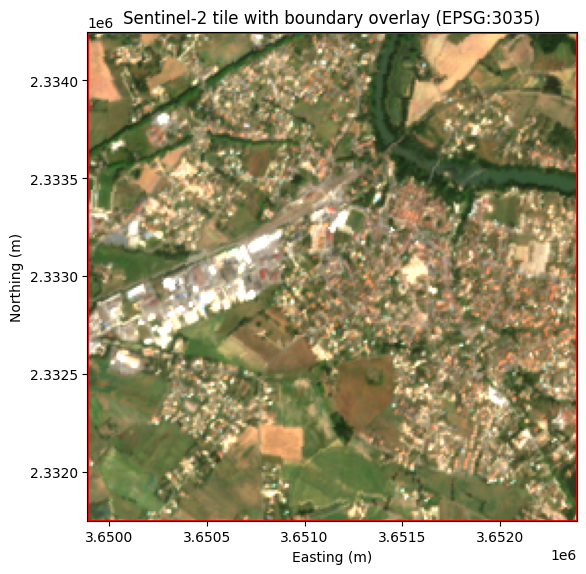
Goal: Create a GeoDataFrame from the tile’s bounding box, then convert it from EPSG:3035 to EPSG:4326 (WGS84). Print the bounds in both CRS.
Steps:
- Create a
shapely.geometry.boxfromtile_bounds - Build a
GeoDataFramewith that geometry andcrs="EPSG:3035" - Convert to EPSG:4326 using
.to_crs() - Print
total_boundsin both CRS
import geopandas as gpd
from shapely.geometry import box
# Step 1: Create a box geometry from tile_bounds
tile_geom = ___ # TODO: box(*tile_bounds)
# Step 2: Build a GeoDataFrame in EPSG:3035
gdf = ___ # TODO: gpd.GeoDataFrame(...)
# Step 3: Convert to WGS84
gdf_wgs84 = ___ # TODO: gdf.to_crs(...)
# Step 4: Print bounds in both CRS
print("EPSG:3035 bounds:", gdf.total_bounds)
print("EPSG:4326 bounds:", gdf_wgs84.total_bounds)box(*tile_bounds)unpacks(left, bottom, right, top)into a rectangular polygon.gpd.GeoDataFrame({"tile": ["FRJ27"]}, geometry=[tile_geom], crs="EPSG:3035")creates the GeoDataFrame..to_crs("EPSG:4326")reprojects to latitude/longitude.
import geopandas as gpd
from shapely.geometry import box
tile_geom = box(*tile_bounds)
gdf = gpd.GeoDataFrame({"tile": ["FRJ27"]}, geometry=[tile_geom], crs="EPSG:3035")
gdf_wgs84 = gdf.to_crs("EPSG:4326")
print("EPSG:3035 bounds:", gdf.total_bounds)
print("EPSG:4326 bounds:", gdf_wgs84.total_bounds)Goal: Identify which NUTS3 region(s) a Sentinel-2 tile intersects using a spatial join, and compute the tile’s area in km².
A spatial join (gpd.sjoin) combines two GeoDataFrames based on the geometric relationship between their features (e.g. intersects, contains, within). This is useful for linking raster tiles to administrative boundaries.
Steps:
- Load NUTS3 boundaries from the Eurostat GeoJSON endpoint (EPSG:3035)
- Create a GeoDataFrame for the tile (reuse
tile_boundsandtile_crsfrom Exercise 3) - Perform a spatial join with
predicate="intersects" - Print the matching NUTS_ID and NUTS_NAME
- Compute the tile area in km² (
tile_geom.area / 1e6)
import geopandas as gpd
from shapely.geometry import box
# Step 7a: Load NUTS3 boundaries (EPSG:3035)
nuts_url = (
"https://gisco-services.ec.europa.eu/distribution/v2/"
"nuts/geojson/NUTS_RG_01M_2021_3035_LEVL_3.geojson"
)
nuts = ___ # TODO: gpd.read_file(nuts_url)
# Step 7b: Create a GeoDataFrame for the tile
tile_geom = box(*tile_bounds)
tile_gdf = ___ # TODO: gpd.GeoDataFrame(..., geometry=[tile_geom], crs=tile_crs)
# Step 7c: Spatial join — find which NUTS3 regions the tile intersects
joined = ___ # TODO: gpd.sjoin(tile_gdf, nuts, predicate="intersects")
# Step 7d: Print matching regions
for _, row in joined.iterrows():
print(f"NUTS_ID: {row['NUTS_ID']}, NUTS_NAME: {row['NUTS_NAME']}")
# Step 7e: Compute tile area in km²
area_km2 = ___ # TODO: tile_geom.area / 1e6
print(f"Tile area: {area_km2:.2f} km²")gpd.read_file(url)can read GeoJSON directly from a URL.- Build the tile GeoDataFrame the same way as in Exercise 6:
gpd.GeoDataFrame({"tile": ["FRJ27"]}, geometry=[box(*tile_bounds)], crs=tile_crs). gpd.sjoin(left, right, predicate="intersects")returns rows fromleftenriched with columns fromrightwhere geometries intersect.- Since
tile_crsis EPSG:3035 (units in metres),tile_geom.areagives square metres — divide by 1 000 000 for km².
import geopandas as gpd
from shapely.geometry import box
nuts_url = (
"https://gisco-services.ec.europa.eu/distribution/v2/"
"nuts/geojson/NUTS_RG_01M_2021_3035_LEVL_3.geojson"
)
nuts = gpd.read_file(nuts_url)
tile_geom = box(*tile_bounds)
tile_gdf = gpd.GeoDataFrame(
{"tile": ["FRJ27"]}, geometry=[tile_geom], crs=tile_crs
)
joined = gpd.sjoin(tile_gdf, nuts, predicate="intersects")
for _, row in joined.iterrows():
print(f"NUTS_ID: {row['NUTS_ID']}, NUTS_NAME: {row['NUTS_NAME']}")
area_km2 = tile_geom.area / 1e6
print(f"Tile area: {area_km2:.2f} km²")4 Displaying satellite images on a map
4.1 Static map with matplotlib
To display the satellite image in geographic coordinates (latitude/longitude), reproject the bounds to WGS84 using rasterio.warp.transform_bounds:
from rasterio.warp import transform_bounds
west, south, east, north = transform_bounds(
tile_crs, "EPSG:4326", *tile_bounds
)
print(f"WGS84 extent: W={west:.4f}, S={south:.4f}, E={east:.4f}, N={north:.4f}")
fig, ax = plt.subplots(figsize=(6, 6))
ax.imshow(rgb, extent=[west, east, south, north])
ax.set_xlabel("Longitude")
ax.set_ylabel("Latitude")
ax.set_title("Sentinel-2 tile in WGS84 coordinates")
plt.tight_layout()
plt.show()WGS84 extent: W=1.6639, S=43.7589, E=1.6979, N=43.7838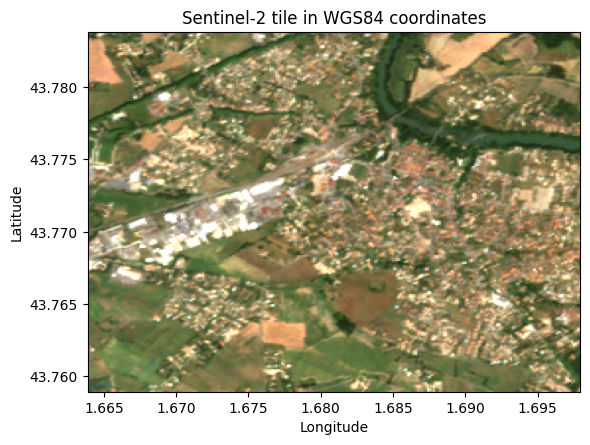
4.2 Interactive map with folium
For an interactive web map, use Folium to overlay the satellite image on a basemap:
import folium
from folium.raster_layers import ImageOverlay
center_lat = (south + north) / 2
center_lon = (west + east) / 2
m = folium.Map(location=[center_lat, center_lon], zoom_start=14)
ImageOverlay(
image=rgb,
bounds=[[south, west], [north, east]],
opacity=0.7,
).add_to(m)
mGoal: Reproject the tile bounds to WGS84 and display the RGB composite on an interactive folium map.
Steps:
- Use
transform_bounds()to convert the tile bounds from EPSG:3035 to EPSG:4326 - Compute the centre of the tile (average of south/north and west/east)
- Create a
folium.Mapcentred on the tile - Add an
ImageOverlaywith the RGB array and WGS84 bounds
import folium
from folium.raster_layers import ImageOverlay
from rasterio.warp import transform_bounds
# Step 1: Reproject bounds to WGS84
west, south, east, north = ___ # TODO: transform_bounds(...)
# Step 2: Compute the centre
center_lat = ___ # TODO
center_lon = ___ # TODO
# Step 3: Create the folium map
m = ___ # TODO: folium.Map(...)
# Step 4: Add the satellite image overlay
___ # TODO: ImageOverlay(...).add_to(m)
mtransform_bounds(tile_crs, "EPSG:4326", *tile_bounds)returns(west, south, east, north)in degrees.- Centre:
center_lat = (south + north) / 2,center_lon = (west + east) / 2. folium.Map(location=[center_lat, center_lon], zoom_start=14)creates the map.ImageOverlay(image=rgb, bounds=[[south, west], [north, east]], opacity=0.7).add_to(m)overlays the image.
import folium
from folium.raster_layers import ImageOverlay
from rasterio.warp import transform_bounds
west, south, east, north = transform_bounds(
tile_crs, "EPSG:4326", *tile_bounds
)
center_lat = (south + north) / 2
center_lon = (west + east) / 2
m = folium.Map(location=[center_lat, center_lon], zoom_start=14)
ImageOverlay(
image=rgb,
bounds=[[south, west], [north, east]],
opacity=0.7,
).add_to(m)
m5 CLC+ Backbone Labels
CLC+ Backbone is a high-resolution land-cover product produced by the Copernicus Land Monitoring Service. It provides pixel-level land-cover classification across Europe at 10 m resolution, making it an ideal ground-truth dataset for training and evaluating segmentation models.
5.1 Land-cover classes
CLC+ Backbone uses 10 land-cover classes:
| Class ID | Name | Description |
|---|---|---|
| 1 | Sealed | Buildings, roads, paved surfaces |
| 2 | Woody – needle leaved | Coniferous forests |
| 3 | Woody – broadleaved deciduous | Deciduous forests |
| 4 | Woody – broadleaved evergreen | Evergreen broadleaved forests |
| 5 | Low-growing woody plants | Bushes, shrubs |
| 6 | Permanent herbaceous | Grasslands, meadows |
| 7 | Periodically herbaceous | Croplands |
| 8 | Lichens and mosses | Natural low vegetation |
| 9 | Non- and sparsely-vegetated | Bare soil, rock |
| 10 | Water | Rivers, lakes, sea |
5.2 Downloading labels
Labels can be downloaded through the Copernicus Land Monitoring Service REST API. The download_clcpluslabel() utility function handles this:
from utils import download_clcpluslabel, tiff_to_numpy
# Bounding box matching our FRJ27 tile (EPSG:3035)
bbox = [3649890, 2331750, 3652390, 2334250]
download_clcpluslabel("label_2018.tif", bbox, year=2018)
label_array = tiff_to_numpy("label_2018.tif") # 2D array of class IDs (1-10)The function queries the WCS endpoint of the CLC+ Backbone service, requesting the raster data for the specified bounding box and year.
5.3 Label–image alignment
The CLC+ labels and Sentinel-2 patches share the same CRS (EPSG:3035) and bounding box, which means they are spatially aligned by construction. Each pixel in the label corresponds to the same geographic area as the matching pixel in the satellite image.
Goal: Download the CLC+ Backbone land-cover label that matches the FRJ27 Sentinel-2 patch and convert it to a NumPy array.
Steps:
- Define the bounding box matching the FRJ27 tile (EPSG:3035):
[3649890, 2331750, 3652390, 2334250] - Call
download_clcpluslabel()with the filename, bounding box, and year - Convert the downloaded TIFF to a NumPy array using
tiff_to_numpy()
from utils import download_clcpluslabel, tiff_to_numpy
# Bounding box matching the FRJ27 tile (EPSG:3035)
bbox = [3649890, 2331750, 3652390, 2334250]
year = 2018
filename = "test_label.tif"
# Step 1: Download the label
___ # TODO
# Step 2: Convert to NumPy array
img_array = ___ # TODOdownload_clcpluslabel(filename, bbox, year)downloads the label and saves it as a TIFF file.tiff_to_numpy(filename)opens the TIFF, converts it to a NumPy array, cleans up nodata values (254, 255 → 0), and returns the array.
from utils import download_clcpluslabel, tiff_to_numpy
bbox = [3649890, 2331750, 3652390, 2334250]
year = 2018
filename = "test_label.tif"
download_clcpluslabel(filename, bbox, year)
img_array = tiff_to_numpy(filename)The training pipeline in section 2 expects labels as NumPy .npy files stored on S3, following a specific path convention. The expected S3 path structure is:
s3://projet-hackathon-ntts-2025/data-preprocessed/labels/CLCplus-Backbone/SENTINEL2/{NUTS}/{year}/{tile_size}/{patch_id}.npy.npy and upload to S3
Goal: Save the label array from Exercise 9 as a .npy file and upload it to S3 in the path expected by the training pipeline.
Steps:
- Save
img_arraylocally as a.npyfile usingnp.save() - Initialise an
S3FileSystemwithget_file_system()(fromutils, same as Exercise 2) - Build the S3 destination path matching the training pipeline convention
- Upload the local file with
fs.put() - Verify the upload by listing the S3 directory with
fs.ls()
import numpy as np
from utils import get_file_system
# Step 1: Save the label array locally as .npy
local_path = "3649890_2331750_0_937.npy"
___ # TODO: np.save(local_path, img_array)
# Step 2: Initialise the S3 filesystem
fs = ___ # TODO: get_file_system()
# Step 3: Build the S3 destination path
s3_path = ___ # TODO: "projet-hackathon-ntts-2025/data-preprocessed/labels/CLCplus-Backbone/SENTINEL2/FRJ27/2018/250/3649890_2331750_0_937.npy"
# Step 4: Upload to S3
___ # TODO: fs.put(local_path, s3_path)
# Step 5: Verify the upload
print(fs.ls(___)) # TODO: S3 directory path (without filename)np.save("file.npy", array)saves a NumPy array to disk.get_file_system()returns ans3fs.S3FileSystemconfigured with the SSP Cloud credentials.- The S3 path must not include the
s3://prefix when usingfs.put()andfs.ls(). fs.put(local_path, s3_path)uploads a local file to S3.fs.ls("projet-hackathon-ntts-2025/data-preprocessed/labels/CLCplus-Backbone/SENTINEL2/FRJ27/2018/250")lists all files in the directory.
import numpy as np
from utils import get_file_system
local_path = "3649890_2331750_0_937.npy"
np.save(local_path, img_array)
fs = get_file_system()
s3_path = "projet-hackathon-ntts-2025/data-preprocessed/labels/CLCplus-Backbone/SENTINEL2/FRJ27/2018/250/3649890_2331750_0_937.npy"
fs.put(local_path, s3_path)
print(fs.ls("projet-hackathon-ntts-2025/data-preprocessed/labels/CLCplus-Backbone/SENTINEL2/FRJ27/2018/250"))Goal: Create a figure with two subplots: the Sentinel-2 RGB composite on the left and the CLC+ label map on the right, with a shared legend showing all 10 land-cover classes.
Steps:
- Create a figure with
plt.subplots(1, 2, ...) - Display the RGB array on the left subplot
- Display the label array on the right subplot with the CLC+ colormap
- Add a legend describing the 10 land-cover classes using
Patch
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from matplotlib.patches import Patch
# Classes & colormap (provided)
classes = [
("Sealed (1)", "#FF0100"),
("Woody -- needle leaved trees (2)", "#238B23"),
("Woody -- Broadleaved deciduous trees (3)", "#80FF00"),
("Woody -- Broadleaved evergreen trees (4)", "#00FF00"),
("Low-growing woody plants (bushes, shrubs) (5)", "#804000"),
("Permanent herbaceous (6)", "#CCF24E"),
("Periodically herbaceous (7)", "#FEFF80"),
("Lichens and mosses (8)", "#FF81FF"),
("Non- and sparsely-vegetated (9)", "#BFBFBF"),
("Water (10)", "#0080FF"),
]
cmap = ListedColormap([color for _, color in classes])
# Step 1: Create the figure
fig, axes = ___ # TODO: plt.subplots(1, 2, figsize=(12, 6))
# Step 2: Sentinel-2 RGB on the left
___ # TODO: axes[0].imshow(rgb)
___ # TODO: set title and turn off axis
# Step 3: CLC+ label on the right
___ # TODO: axes[1].imshow(img_array, cmap=cmap, vmin=1, vmax=10)
___ # TODO: set title and turn off axis
# Step 4: Legend
legend_elements = [
___ # TODO: Patch(facecolor=color, edgecolor="black", label=label)
# for label, color in classes
]
fig.legend(
handles=legend_elements,
loc="center right",
bbox_to_anchor=(1.35, 0.5),
frameon=True,
)
plt.tight_layout()
plt.show()plt.subplots(1, 2, figsize=(12, 6))returns a figure and an array of two axes.- Use
axes[0].imshow(rgb)for the satellite image, andaxes[1].imshow(img_array, cmap=cmap, vmin=1, vmax=10)for the label. - Turn off axes ticks with
axes[i].axis("off"). - Build the legend with a list comprehension:
[Patch(facecolor=color, edgecolor="black", label=label) for label, color in classes].
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from matplotlib.patches import Patch
classes = [
("Sealed (1)", "#FF0100"),
("Woody -- needle leaved trees (2)", "#238B23"),
("Woody -- Broadleaved deciduous trees (3)", "#80FF00"),
("Woody -- Broadleaved evergreen trees (4)", "#00FF00"),
("Low-growing woody plants (bushes, shrubs) (5)", "#804000"),
("Permanent herbaceous (6)", "#CCF24E"),
("Periodically herbaceous (7)", "#FEFF80"),
("Lichens and mosses (8)", "#FF81FF"),
("Non- and sparsely-vegetated (9)", "#BFBFBF"),
("Water (10)", "#0080FF"),
]
cmap = ListedColormap([color for _, color in classes])
fig, axes = plt.subplots(1, 2, figsize=(12, 6))
axes[0].imshow(rgb)
axes[0].set_title("Sentinel-2 RGB")
axes[0].axis("off")
axes[1].imshow(img_array, cmap=cmap, vmin=1, vmax=10)
axes[1].set_title("CLC+ Label")
axes[1].axis("off")
legend_elements = [
Patch(facecolor=color, edgecolor="black", label=label)
for label, color in classes
]
fig.legend(
handles=legend_elements,
loc="center right",
bbox_to_anchor=(1.35, 0.5),
frameon=True,
)
plt.tight_layout()
plt.show()